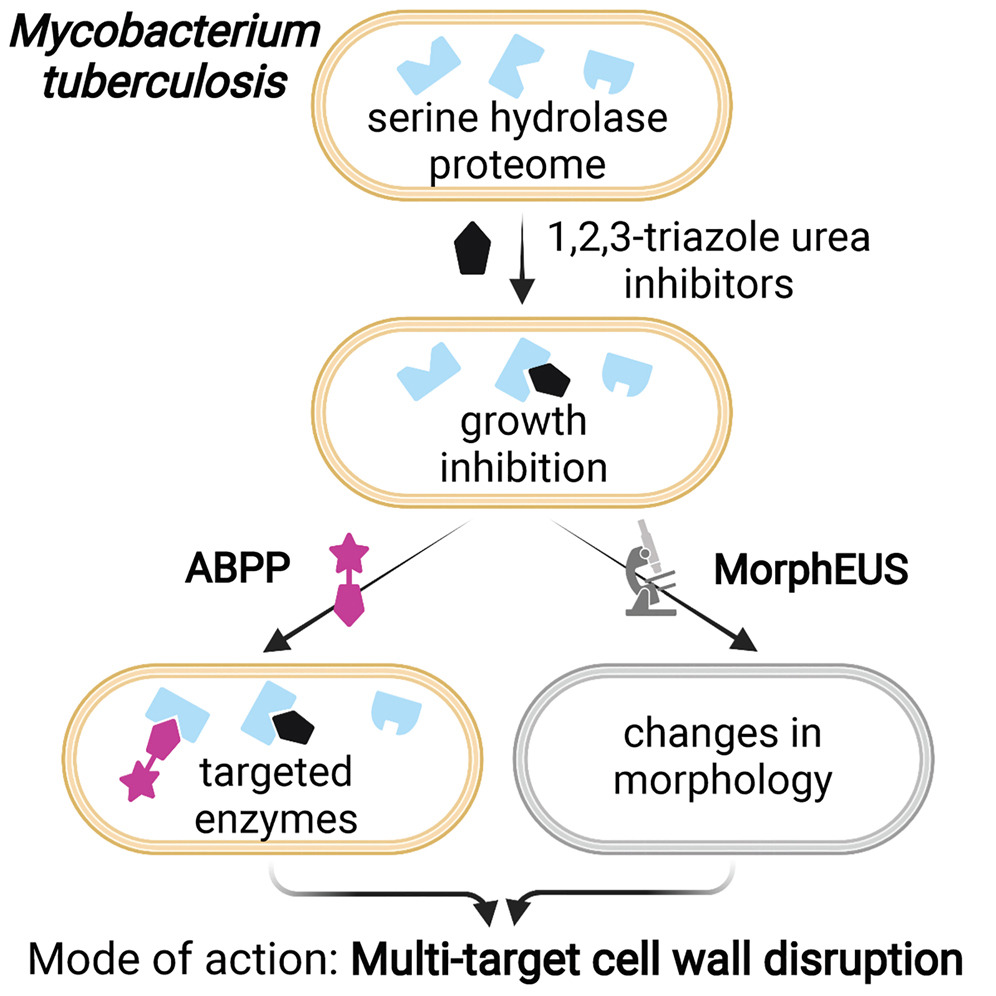
The identification and validation of a small molecule’s targets is a major bottleneck in the discovery process for tuberculosis antibiotics. Activity-based protein profiling (ABPP) is an efficient tool for determining a small molecule’s targets within complex proteomes. However, how target inhibition relates to biological activity is often left unexplored. Here, we study the effects of 1,2,3-triazole ureas on Mycobacterium tuberculosis (Mtb). After screening ∼200 compounds, we focus on 4 compounds that form a structure-activity series. The compound with negligible activity reveals targets, the inhibition of which is functionally less relevant for Mtb growth and viability, an aspect not addressed in other ABPP studies. Biochemistry, computational docking, and morphological analysis confirms that active compounds preferentially inhibit serine hydrolases with cell wall and lipid metabolism functions and that disruption of the cell wall underlies biological activity. Our findings show that ABPP identifies the targets most likely relevant to a compound’s antibacterial activity.
By Li, M.; Patel, H. V.; Cognetta, A. B.; Smith, T. C.; Mallick, I.; Cavalier, J. F.; Previti, M. L.; Canaan, S.; Aldridge, B. B.; Cravatt, B. F.; Seeliger, J. C
Full paper here
Published on 06/10/2021






